Translocation t(12;16) (q13;p11) is found in approximately 95 % of myxoid liposarcomas.
It leads to a joining of the genes TLS (FUS) on the chromosome 16 (exone/introne structure homologous with EWS) with the gene CHOP (DDIT3, GADD153) from the chromosome 12. The points of break are relatively variable. Breakages are most often found in introns 5, 7 and 8 of the gene TLS and in the introne 1 of the gene CHOP. Various types of joinings then give rise to the various types of gene fusions (Type 1 – 4). The fusion gene encodes a chimeric protein, where N-terminal region of the TLS gene with activating domain is fused with the complete gene CHOP, the transcription factor from bZIP family.
Note. part of the myxoid liposarcomas carry the translocation t(12;22) (q13;q12) EWS/CHOP.
Examination
The presence of translocation t(12;16), or the CHOP gene breakage is analyzed using RT-PCR and FISH.
RT-PCR
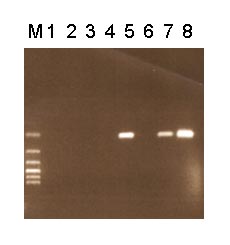
When RT-PRC, we isolate mRNA and after its transcription into cDNA we detect chimeric transcripts using PCR. PCR products are vizualized through agarose gel electrophoresis (fig. 1). (Because of the size of the PCR product type 1 (379 bp) it is necessary to perform the analysis from fresh material)
FISH
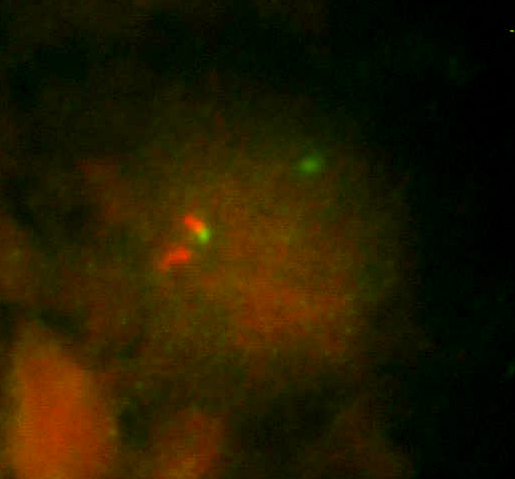
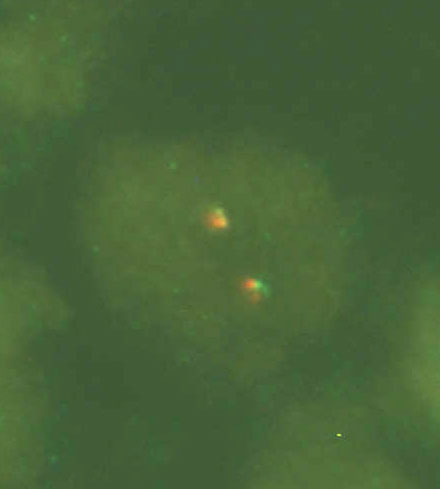
When FISH, we detect the break of the gene CHOP using CHOP Dual Color, Break Apart Rearrangement Probe, Vysis/Abbot (this probe does not identify the specific translocation partner). In a sample positive for rearrangement of the gene CHOP we observe one yellow, one red and one green signal (fig. 2A). In a negative sample we observe two yellow signals (fig. 2B). (dual color filter)
References
- Rabbitts TH, Forster A, Larson R, Nathan P. Fusion of the dominant negative transcription regulator CHOP with a novel gene FUS by translocation t(12;16) in malignant liposarcoma. Nat Genet. 1993;4(2):175-180.
- Adams V, Hany MA, Schmid M, Hassam S, Briner J, Niggli FK. Detection of t(11;22)(q24;q12) translocation breakpoint in paraffin-embedded tissue of the Ewing's sarcoma family by nested reverse transcription-polymerase chain reaction. Diagn Mol Pathol. 1996 Jun;5(2):107-13.
- Hisaoka M, Tsuji S, Morimitsu Y, Hashimoto H, Shimajiri S, Komiya S, Ushijima M. Detection of TLS/FUS-CHOP fusion transcripts in myxoid and round cell liposarcomas by nested reverse transcription-polymerase chain reaction using archival paraffin-embedded tissues.Diagn Mol Pathol. 1998 Apr;7(2):96-101.
- Hill DA, O'Sullivan MJ, Zhu X, Vollmer RT, Humphrey PA, Dehner LP, Pfeifer JD.Practical application of molecular genetic testing as an aid to the surgical pathologic diagnosis of sarcomas: a prospective study.Am J Surg Pathol. 2002;26(8):965-977
- Pfeifer JD (Ed). Molecular Genetic Testing in Surgical Patology. LWW 2006. Philadelphia, USA.