Monoclonality is considered to be the essential characteristic of neoplasia. Detection of mono or polyclonality can therefore play a significant role in the differentiation of some malignant neoplasia which are mostly monoclonal from polyclonal reactive lesions. There have been developed many methodologies for clonality analysis. One of them is the method which uses the phenomenon called lyonization, i.e. random inactivation of the X chromosome caused by methylation in individuals with a set of chromosomes XX. The theory says that to compensate a gene dosage during the early embryogenesis, one of the two parental copies of X chromosome is inactivated by methylation in each cell. It is supposed that this methylation is irreversible and methylated paternal or maternal chromozome is inherited by all daughter cells. The ratio of the methylated X chromosomes received from the mother or father in the majority of a normal female tissue is about the same. On the contrary, in neoplasia where the population of the cells arised from one mother tumor cell, and so the same one chromosome remains methylated, there occurs the inclination of previous average towards the maternal or paternal X chromosome. One of the analysis which detects this imbalance is also a method using the HUMARA locus and methylation-sensitive restriction enzymes.
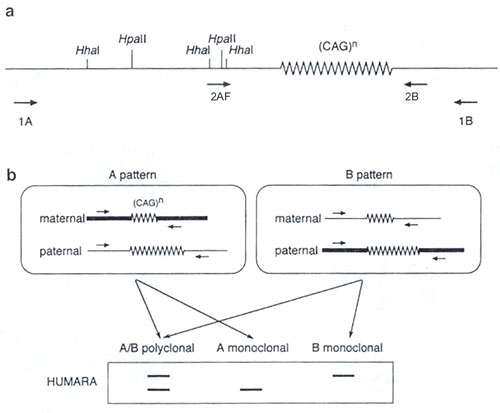
Locus for the human androgen receptor (HUMARA) is located on the chromosome X in the region q13. His first exon contains trinucleotide (CAG) short tandem repeat (STR) with a relatively considerable variability (heterozygosity about 90 %). This STR lies close to the 5´ promotor of the HUMARA gene which contains CpG dinucleotides (CpG islands) in a high frequency. The test uses polymerase chain reaction (PCR), high variability of the HUMARA locus and specific properties of the restriction endonucleases Hhal and Hpall whose restriction sites are located in CpG islands, i.e. inside the potential amplicon. Before PCR, cleavage of the investigated DNA using these enzymes is performed. Thanks to their sensitivity to methylated DNA only the unmethylated chromosome is cleaved. Thus during PCR there occurs only the amplification of uncleaved, i.e. methylated sequences. Then, by the separation of cleaved and uncleaved amplicons on acrylamide gel, and by their comparison, we can distinguish monoclonal and polyclonal status of the sample (Fig. 1).
Examination
Clonality analysis using the testing of X chromosome methylation status in the HUMARA locus begins by the cleavage of DNA using the methylation-sensitive restriction enzymes (HhaI and HpaII).
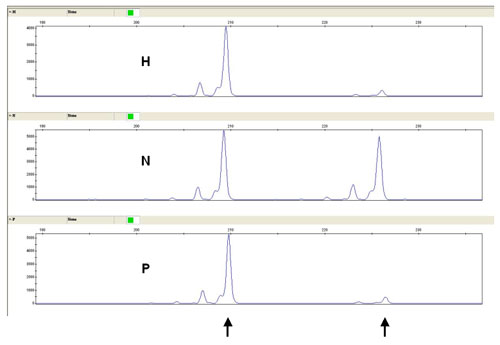
This is followed by PCR of cleaved and uncleaved DNA which amplifies STR HUMARA. There is used a set of primers labeled with fluorescent substance 6-FAM. PCR products are then separated using an automatic sequencer ABI Prism 3130 XL. Fragmentation analysis compares the pattern of cleaved and uncleaved samples (Fig. 2)
References
- Vogelstein B, Fearon ER, Hamilton SR, Feinberg AP. Use of restriction fragment length polymorphisms to determine the clonal origin of human tumors. Science. 1985;227(4687):642-5.
- Allen RC, Zoghbi HY, Moseley AB, Rosenblatt HM, Belmont JW. Methylation of HpaII and HhaI sites near the polymorphic CAG repeat in the human androgen-receptor gene correlates with X chromosome inactivation. Am J Hum Genet. 1992;51(6):1229-39.
- Paradis V, Laurent A, Flejou JF, Vidaud M, Bedossa P. Evidence for the polyclonal nature of focal nodular hyperplasia of the liver by the study of X-chromosome inactivation. Hepatology. 1997;26(4):891-5.
- Nakamura H, Hirota S, Adachi S, Ozaki K, Asada H, Kitamura Y. Clonal nature of seborrheic keratosis demonstrated by using the polymorphism of the human androgen receptor locus as a marker. J Invest Dermatol. 2001;116(4):506-10.
- el Hamidi A, Kocjan G, Du MQ. Clonality analysis of archival cervical smears. Correlation of monoclonality with grade and clinical behavior of cervical intraepithelial neoplasia. Acta Cytol. 2003;47(2):117-23.